Learning Recurrent Waveforms Within EEGs
Austin J. Brockmeier and Jose C. Principe
IEEE Trans. on Biomedical Engineering,
Vol. 63,
No. 1,
pp. 43-54,
2016
Overview
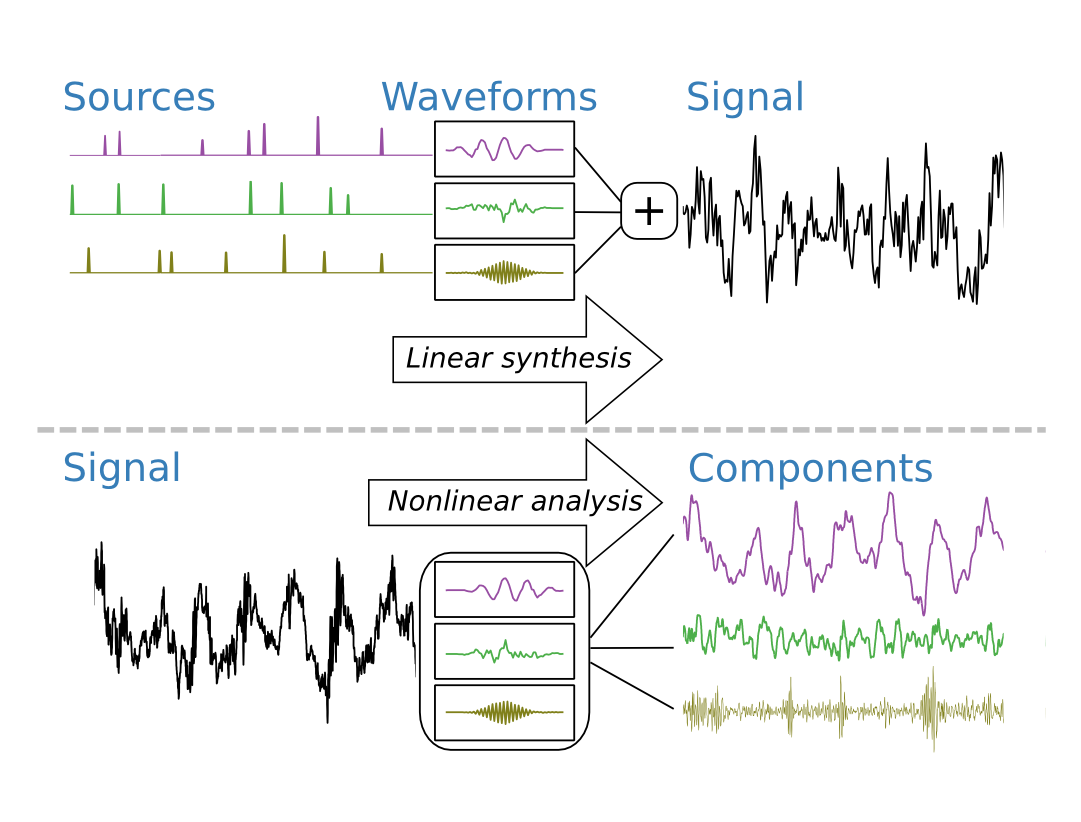
When experts analyze EEGs they look for landmarks in the traces corresponding to established patterns such as oscillatory and phasic events of particular frequency or morphology. Long records motivate automated analysis techniques. Automation techniques often require design choices such as wavelet family or number of bandpass filters. To overcome this, we explore a modeling approach that automatically learns recurrent temporal waveforms within EEG traces. The estimation is based on a multiple-input, single-output linear model with sparsely excited inputs.
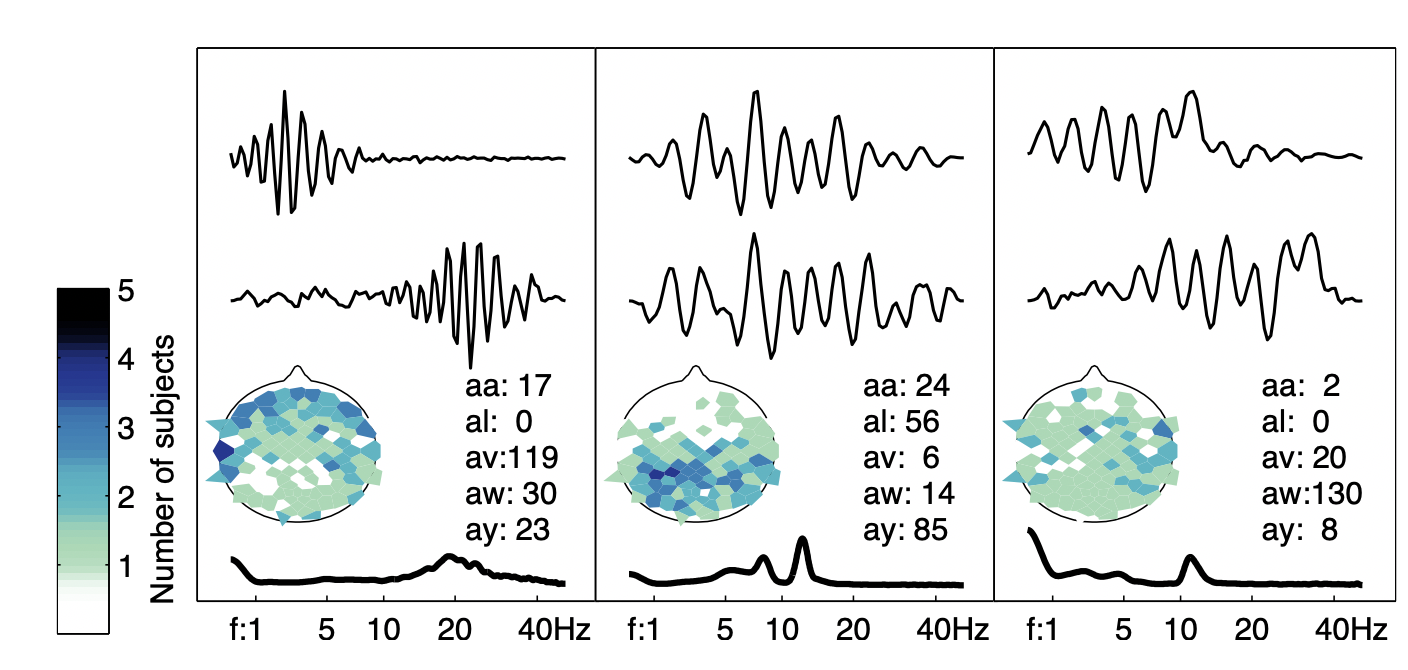
We apply the approach to various data sets to better understand the specificity and consistency of the estimated waveforms. In particular, we cluster the waveforms found on different electrodes/channels and subjects.