Joint Penn-UDel Colloquium on the Nature of Computing
Physical Mapping of DNA Fragments Using Genetic Algorithms
Walter Cedeńo

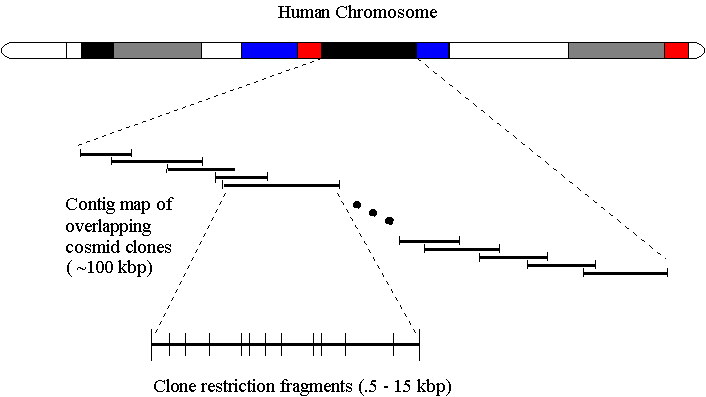
The goal of the Human Genome Center at Lawrence Livermore National Laboratory (LLNL) is to construct a physical map of chromosome 19. The physical map in conjunction with the sequence information allows researchers to study genome organization and variation. Constructing the physical map of chromosome 19 is an enormous task. Using restriction enzymes and cloning techniques the chromosome is divided into overlapping cosmid clones composed of a set of restriction fragments. During the process, the order of both the fragments in the clones and the clones in the chromosome is lost.
Given a set of clones with their restriction-fragment data the objective is
to reconstruct the order of the sequence of clones in the DNA. This problem
is similar to one of putting together a linear jigsaw puzzle and is known to
be NP-hard. Here we present a Genetic Algorithm (GA) approach for
reconstructing the overlapping sequence of clones. The GA is tested using
different data sets obtained from the Human Genome Project at LLNL. Results
will be presented for our GA approach and that of other more classical GAs
techniques.
Friday, November 20, at 4 pm
in Room 207 of the
Johnson Pavilion
(see map)
near Spruce and 36th Streets on the
University of Pennsylvania campus.
Travel Directions are found at
http://www.upenn.edu/fm/map/dir.html